+1 Level Up
The world’s only single-cell DNA sequencing technology platform gets an upgrade
Jan Cools is Group Leader at the Vlaams Instituut voor Biotechnologie (VIB) Center for Cancer Biology in Leuven, Belgium. We spoke with him about his experience using Mission Bio’s recently upgraded single-cell DNA sequencing platform, Tapestri, and how it could speed up new research in the field.
What’s your background?
I started my journey as a scientist in the last year of my studies at the University of Leuven (KU Leuven) in 1996, where I immediately became involved in a project to clone the JAK2 gene and determine its exon-intron structure. Back then, the human genome sequence was not yet available!
It was an amazing time to be performing genetic studies in hematology and oncology; many new oncogenes were identified and many new mutations discovered. During my PhD and postdoc (at KU Leuven and Harvard Medical School, respectively), we worked on leukemias with aberrant karyotypes to identify the oncogenes involved using basic molecular methods such as PCR. Later, we added array-CGH to our approaches and, later still, next-generation sequencing, which changed the field completely. Now that we can carry out single-cell sequencing in thousands of cells at the RNA or DNA level, the field is once again taking major steps forward.
Why is single-cell RNA sequencing important to your work?
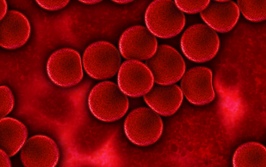
Single-cell RNA sequencing is a powerful tool for identifying all the cell types and states present in a complex sample, such as the blood or bone marrow of a leukemia patient – but it cannot directly distinguish between wild-type and mutant cells. Mission Bio’s single-cell DNA sequencing technology allows scientists like me to identify the point mutations, indels, and copy number changes in the DNA of each cell, providing clear information on how many mutations are present in each of the 6,000 to 8,000 cells analyzed for each sample.
How did you first encounter the Tapestri platform?
At the VIB Institute in Leuven, we have a “TechWatch” team who look for new technologies that could be of interest to VIB researchers. When the team identified the Tapestri platform early on, I was immediately interested in using it to analyze the heterogeneity of leukemia samples. The platform’s recent update allows researchers to work with fewer cells (20,000, down from a minimum of 100,000), includes a number of new ready-to-use gene panels (such as the panel we designed for acute lymphoblastic leukemia and others in the hematology-oncology field), and is further improved for simultaneous single-cell DNA and protein analysis.
The new ready-to-use panels will help kickstart experiments by removing the need to first design and test all the PCR fragments needed to sequence the DNA. These new panels have been pre-designed and tested; everyone can see which genes are included and decide which panel is best for their purpose. The addition of antibodies to the samples allows researchers like me to identify surface markers on the cells they are sequencing and obtain DNA sequence data linked to protein expression, helping to identify the types of cells harboring specific mutations.
Which patients are you hoping the upgrade will benefit?
There are now gene panels available for most hematological cancers, which should boost the entire heme/onc field.
Between studying for my English undergrad and Publishing master's degrees I was out in Shanghai, teaching, learning, and getting extremely lost. Now I'm expanding my mind down a rather different rabbit hole: the pharmaceutical industry. Outside of this job I read mountains of fiction and philosophy, and I must say, it's very hard to tell who's sharper: the literati, or the medicine makers.